Scientists find a golden way to detect harmful bacteria
Ashley Roby
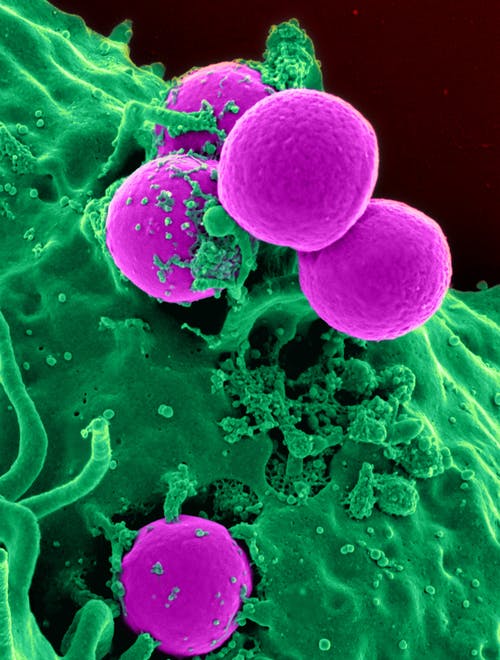 Credits: Pixabay
Credits: Pixabay
This article talks about a new method scientists have devised to detect bacterial contamination in food and water samples - using gold nanoparticles!
Tweet
Harmful bacteria like Staphylococcus and Pseudomonas can grow inside our body and cause various diseases. The species of Pseudomonas called Pseudomonas aeruginosa is especially terrible because it can resist multiple drugs and cause long-term diseases. It can infect patients with burn injuries and weakened immune systems, thus earning the title - opportunistic pathogen. It has remained a leading cause of hospital-acquired infections. According to the CDC (Centre for Disease Control and Prevention), in 2017, Pseudomonas caused 32,600 infections in hospitalised patients and 2,700 deaths in the US. It became the sixth most frequently occurring pathogen and the second most common cause of ventilator-associated pneumonia. Pseudomonas mainly makes its way through contaminated objects, food, or water samples.
This grave scenario demands rapid diagnostic techniques to safeguard the patients and contain a possible hospital outbreak. However, existing methods of testing samples still rely on bacterial culturing, a method of growing bacteria on a culture medium that selectively promotes its growth. However, this method may take several days and give false results due to improper handling of the sample. Even though a few other modern techniques like Real-Time Polymerase chain reaction (RT-PCR) that detect RNA in the sample can provide accurate and sensitive results, they require expensive equipment and trained personnel - becoming unsuitable for cheaper diagnosis.
A group of researchers from Jiangnan University, China, recently discovered a simple, quick, and cheap method to detect these bacteria from contaminated samples. One can obtain results using a simple portable scanner in just under 20 minutes. This technique involves a gold nanoparticle-based immunochromatographic assay (ICA) strip that gives an easily readable colour change. However, samples containing few bacteria need to undergo bacterial enrichment using favourable growth media so that they can be detected. The enrichment process can take up to 8 hours, and a quick ICA strip test will give the result in 15-20 minutes. This is a faster detection method as compared to standard bacterial culturing that takes up to 2-3 days.
 Credits: CDC, Unsplash
Credits: CDC, Unsplash
The detection method uses two things - antibodies or proteins that specifically bind on to the surface of Pseudomonas and gold nanoparticles, which have an intense red color The nanoparticles are attached to these antibodies and mixed with a sample. When one applies the mixture to an ICA strip, it can give two kinds of signals. The strip will show a test signal and a control signal for a sample contaminated with Pseudomonas. In contrast, it will show only a control signal from a contamination-free sample. The control helps us know if the strip is working correctly or not, and the intensity of the test signal tells us the amount of contamination in the sample.
 Credits: Mostera, Pexels
Credits: Mostera, Pexels
The strip proves advantageous because it does not give signals for other commonly found bacteria in water samples. Hence, the group of researchers succeeded in developing a technique that produces an objective and easily interpretable output in a short time.
Across the globe, doctors have reported high rates of infections caused by drug-resistant Pseudomonas aeruginosa. The morbidity and mortality rates associated with this strain range from 18%-61%, making it a public health threat that needs immediate attention. The development of this ICA strip test technique is an efficient headstart to check for Pseudomonas contamination in food and water samples. Early detection of this bacteria can help prevent infection, spread, and save millions of lives. This study also lays a good foundation for future research to develop antibodies that are targeted explicitly against Pseudomonas.
Bibliography
A voracious reader and science enthusiast, Ashley is forever trying (and failing) to balance her interests and manage 6 hours of sleep. She is currently pursuing a BS-MS degree from IISER TVM and looks forward to becoming a science journalist and a fiction writer in the future. (who says you can’t be both?)
Note: This article was submitted by Ashley Roby as an assignment during the workshop Scicomm for Scientists 2021, organised by Cogito137, IISER Kolkata, funded by the Department of Science and Technology, Govt. of India. The assignment was selected for publication and has undergone due editorial process. Team Cogito137 thanks Spoorthy Raman for the initial editorial review of this article.
signup with your email to get the latest articles instantly